Biological Sciences
Applications for 2024-2025 open 1 July 2024
Novel antibiotics from the genomic dark matter
Project code: SCI001
Supervisor:
Dr Stephanie Stuteley
Discipline: School of Biological Sciences
Project description
Naturally occurring antibiotics effective against human pathogens are of medical and commercial interest. These important molecules, known for their complex structures, are produced by biosynthetic enzymes. This project aims to characterise enzymes identified from metagenomic sequences that catalyse the formation of molecules with potential antimicrobial properties. The purified enzymes will be used to produce mature and biologically active compounds in vitro.
Students with an interest in biochemistry, molecular biology or structural biology are encouraged to apply. During this project, students will gain hands-on experience in protein expression, purification and biochemical assays.
Regulation of central carbon metabolism in pathogenic bacteria
Project code: SCI002
Supervisor:
Dr Jamie Taka
Discipline: School of Biological Sciences
Project description
Mycobacterium tuberculosis (Mtb), the causative agent of tuberculosis, is the leading cause of death worldwide from a single infectious agent. Isocitrate lyase (ICL) enzymes enable Mtb to efficiently use lipids as a carbon source during chronic infection, playing a crucial in the persistence, virulence, and antibiotic tolerance of Mtb. This project aims to investigate how ICL enzymes regulate central carbon metabolism in Mtb.
Students with a background in protein science will be suitable for this position. The selected applicant will receive training in protein expression and purification techniques, as well as various biochemical, biophysical, and structural tools.
This project is funded by the Marsden Fund to support applicants who whakapapa Māori.
Korimako (Bellbird): song reproductive success and movement in a fragmented landscape
Project code: SCI003
Supervisor:
Discipline: School of Biological Sciences
Project description
Bellbird are one of New Zealand’s most iconic songbirds and play an important role as pollinators in New Zealand forest ecosystems. This project focusses on populations of bellbird in two mainland sanctuaries (Tawharanui and Shakespear parks). The project aims to understand the role of habitat quality, reproductive success and animal movements within a fragmented landscape. Does habitat quality influence body condition and reproductive success of bellbirds? Are singing performances and song diversity influenced by habitat quality? What influence does forest fragmentation have on bird movements and the formation of local song neighbourhoods? This research is part of a PhD project. The summer student working on this project will learn field and analyses skills and requires a reasonable level of physical ability and interest in the outdoors.

Vocal learning and song development in a translocated population of Tieke Saddleback
Project code: SCI004
Supervisor:
Discipline: School of Biological Sciences
Project description
Tieke/Saddleback are an endemic species of songbird that have been the focus of species translocations aimed at saving this species from extinction. This project will focus on the song development of free-living saddleback populations translocated to predator-free mainland sanctuaries (Tawharanui and Shakespear parks). Behavioural observations and sound recordings of banded birds (adults and juveniles) will be collected in the field and analysed using specialised bioacoustics software. The work is part of a PhD project that aims to measure the rate of cultural song evolution and the effects of translocation, habitat quality, and sociality on song learning. The summer student working on this project will learn field and analyses skill and requires a reasonable level of physical ability and interest in the outdoors.

Plant phenological change in University of Auckland grounds: impacts of climate change?
Project code: SCI005
Supervisor:
Discipline: School of Biological Sciences
Project description
One of the impacts of climate change is hypothesised to be changes in timing of plant reproductive events, e.g. flowering. Such changes can impact plant performance in terms of fruit/seed output, or availability of resources for wildlife. The University of Auckland has archival data (on paper) on the phenology (timing) of plant flowering within the University Grounds. This project would be to both digitize and secure this archival information and compare it to the phenology of the same plants in 2024 (as possible) to determine climate change effects at a local level. Such a study would contribute to use of the University Grounds as a Living Laboratory.

Variation in myrtle rust susceptibility of pōhutukawa in Auckland city
Project code: SCI006
Supervisor:
Discipline: School of Biological Sciences
Project description
Myrtle rust (Austropuccinia psidii) is a virulent plant pathogen of the family Myrtaceae, with the rust now having spread globally and infecting >500 plant species. It arrived in New Zealand in 2017 and is still spreading. Because it is a recent arrival, our understanding of host susceptibility and disease progression is also still developing, both within and amongst species. For pōhutukawa within Auckland, infections of individuals seem to vary by infection date, location, plant size, proximity to roads, and whether the plant was planted or wild grown. Sometimes infected plants and healthy plants occur directly next to each other. This proposed project would build on previous surveys of pōhutukawa within Auckland to score the infection status of trees and a range of potential factors (e.g., size, proximity to roads, and likely origin) that influence susceptibility to infection. These data would then be analysed for infection patterns both within and between years. The goal is to determine factors (host, environment) that determine levels of myrtle rust infection to better understand the infection process and inform management response.

Effects of sleep disruption on daytime performance in captive songbirds
Project code: SCI007
Supervisor:
Discipline: School of Biological Sciences
Project description
Sleep, and the lack of sleep, have important consequences for everyday actions, mood and performance. Though birds sleep in a manner similar to humans, we have a very limited understanding of how important sleep is for birds’ daytime activity and behaviour.
We are conducting a large project with captive zebra finch to understand normal bird sleep behaviour, and the consequences of sleep that is disrupted, such as by noise and light at night.
Birds are housed on campus. The work would include observing and coding bird behaviour after normal and disturbed sleep. You would also assist in care and feeding of the birds.
Characterising essential genes required for chloroplast biogenesis
Project code: SCI008
Supervisor:
Discipline: School of Biological Sciences
Project description
Our group has identified several new proteins which are essential for chloroplast biogenesis. This project will look at trying to characterise their functions. You will use a combination of genetics, cell biology, biochemistry and microscopy techniques to try and determine the molecular and physiological functions of the proteins.
Skilled required: Skills in molecular biology and genetics with an interest in plant development/physiology would be beneficial. BIOSCI papers such as 202, 205, 351 and 326 provide a good background.
Controlling fruit size in kiwifruit: Analysis of CRISPR-edited kiwifruit plants
Project code: SCI009
Supervisor:
Discipline: School of Biological Sciences
Project description
Fruit are essential for our day to day diet. The aim of the project is to contribute to our understanding of the mechanisms that are important for fruit development especially fruit size. Genes potentially involved in the control of fruit size have been mutated using targeted gene editing (CRISPR) in kiwifruit. During the project, the aim is to start characterising the edited plants. Key techniques: Plant molecular biology (Bioinformatics – genotyping - gene expression analysis) and Phenotyping.
Peptides targeting cancer cells
Project code: SCI010
Supervisor:
Assoc Prof Paul Harris
Dr Jiwon Hong
Dr Renata Kowalczyk
Discipline: School of Biological Sciences
Project description
Cancer remains one of the biggest global health issues and many cancers do not respond to the established treatments of surgery, chemotherapy, and radiotherapy, impacting survival rates for aggressive cancers such as pancreatic, liver and lung, brain and oesophagus that have extremely poor 5-year survival rates of around 20%.
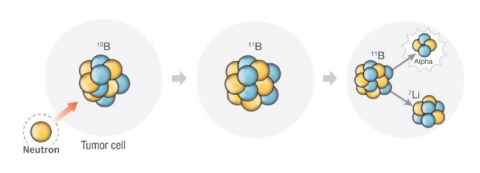
Boron neutron capture therapy (BNCT) is a doubly selective cancer therapy, combining a chemical reagent and a radiation component and has significant advantages compared to chemotherapy and radiotherapy alone. BNCT uses a boron-10 containing compound that accumulates in cancer cells, followed by local irradiation with thermal neutrons. High energy particles are then deposited only within the cancer cell, limiting damage only to cells containing the boron-10 carrier. Revolutionary technology that produces neutron beams via non-nuclear sources is here, but research into boron-10 containing compounds that discriminate for cancer cells has failed to keep pace. Using a specially designed, peptide boron-10 carrier, we can target cancer cells that contain a specific integrin marker, completely absent in healthy cells, thereby delivering a therapeutic load of boron directly to cancerous tissue. Successful candidates will participate in solid phase peptide synthesis to prepare cancer targeting peptides and have an opportunity to study biological assays.
See: Harris et al. Eu. J. Med.Chem 2017, 136, 154. and. Molecules, 2022, 27,4331.
Novel peptide-based antibiotics
Project code: SCI011
Supervisor:
Assoc Prof Paul Harris
Assoc Prof Viji Sarojini
Prof Alan Davidson
Dr Veronika Sander
Discipline: School of Biological Sciences
Project description
Antibiotic resistance has been recognised by the WHO as one of the greatest threats to humanity and infectious diseases rank as the second most common cause of death worldwide. New antibiotics are desperately needed and if nothing is done by 2050 it is estimated > 10M people will die per annum, which is more that cancer and diabetes.
Cyclic lipopeptides are an emerging subset of peptide-based antibiotics (e.g., FDA-approved daptomycin and polymyxin) containing a lipid or fatty acid. They have been shown to possess clinical efficacy and are used as the “last line of defence” against otherwise untreatable bacterial infections. Despite their promise, undesired toxicity is a significant drawback.
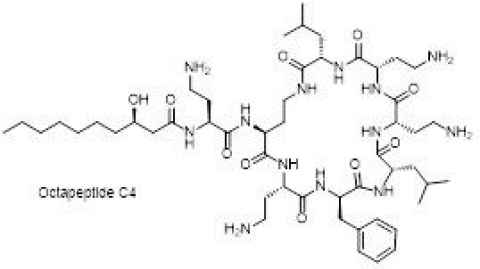
We are developing novel, non-toxic derivatives of naturally occurring lipopeptide antibiotics (e.g., Fig. 1) by modifying the chemistry of the lipid tail. Novel antibiotic analogues will undergo biological testing against multi-drug resistant (MRD) strains of bacteria and evaluation of potential toxicity.
Successful candidates will use organic synthesis and modern methods of solid phase peptide synthesis. Candidates can also undertake and learn biological assays if they desire.
See: Harris et al. ACS Infect. Dis. 2022, 8, 2413
See: Sarojini et al. J. Med. Chem. 2015, 58, 2, 625
Anemone metabolism
Project code: SCI012
Supervisor:
Prof Tony Hickey
Dr Jules Devaux
Discipline: School of Biological Sciences
Project description
Very little is known about cnidarian metabolism (Jelly fish, Hydras, Anemones etc.). Anemones are a useful model to study as we can get them locally and they bud and we have found that they clone in the lab, giving us lots of small anemones to work with. We want to explore how their metabolism works, hardly anything is known about their mitochondria. Not that anemones can occupy rockpools, which are exposed to extreme hypoxia (low oxygen), and some are found locally in sulphurous environments. Hydrogen sulphide typically poisons mitochondria. We aim to see how they may survive these habitats. We have access to respirometry and imaging techniques and possibly magnetic resonance imaging with some NMR spectroscopy.
Measuring milky flesh snapper muscle
Project code: SCI013
Supervisor:
Prof. Tony Hickey
Prof Andrew Jeffs
Dr Darren Parson
Discipline: School of Biological Sciences
Project description
If you have been reading the news you may have seen that a reasonable percentage of snapper (~10%) have been found to have milky flesh, i.e. their swimming muscles have degenerated to a gel like state. The student will help explore this issue by comparing muscle tissues from healthy and pathologically fish samples. The student will learn how to measure enzyme activities using spectroscopy and if available measure respiration of fish muscle.
What does the choanoflagellate genome encode?
Project code: SCI014
Supervisor:
Discipline: School of Biological Sciences
Project description
Choanoflagellates are often overlooked aquatic hunters of bacteria, and the closest living relatives of animals. For the past five years, students in our final year undergraduate protein science paper, BIOSCI350, have been sequencing genes, and producing proteins from the model choanoflagellate Salpingoeca rosetta. We have identified a number of interesting choanoflagellate proteins which we can make in large amount. In this summer project you’ll attempt to isolate and structurally characterize some of these proteins.
Investigating Human Malic Enzymes in cancer: recombinant expression & small molecule screening
Project code: SCI015
Supervisor:
Discipline: School of Biological Sciences
Project description
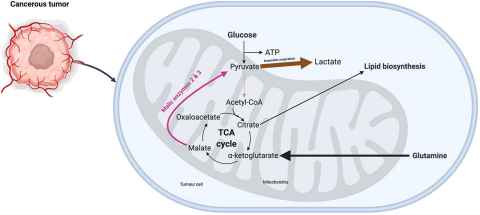
Cancer cells exhibit remarkable adaptability, evading the usual cellular checkpoints to fuel their relentless proliferation. Our research focuses on malic enzyme, a critical player in cancers altered metabolism. By investigating the structural nuances of malic enzyme isoforms, we aim to uncover vulnerabilities that can be exploited for targeted anticancer therapies.
This project will encompass the expression and purification of human malic enzyme isoforms utilizing an existing bacterial expression system. Prior structural elucidation efforts in the Loomes laboratory have facilitated the acquisition of small molecules. These molecules are slated for screening against the three human malic enzyme isoforms to investigate the complex binding interactions with small molecule inhibitors. Contingent upon progress, protein crystallography may commence for any molecules that demonstrate binding affinity.
Ideal student: Skills in protein expression and/or differential scanning fluorimetry (DSF), however, all applicants will be considered!
Searching for new antibiotics
Project code: SCI016
Supervisor:
Assoc Prof Shaun Lott
Dr Stephanie Dawes
Discipline: School of Biological Sciences
Project description
Antibiotics are a special category of drug that underpin modern medicine as we know it. Resistance to antibiotics is a growing global problem, which is estimated will place 10 million lives per year at risk by 2050 without action. This project will focus on identifying inhibitorsthe enzyme RNase HI that is a potential target for new antibiotics against tuberculosis and gonorrhoea. This project is best suited to someone with a strong interest in protein structure and function, medicinal chemistry and/or medical microbiology.
The structure and function of bacterial teneurin-like proteins
Project code: SCI017
Supervisor:
Discipline: School of Biological Sciences
Project description
Our previous work published in Nature (tinyurl.com/lottlab) showed that bacterial RHS/YD repeat sequences encapsulate toxic proteins prior to delivery. However, in the eukaryotic teneurin proteins, the same repeat sequences help form intercellular connections. The bacterial homologues of teneurins are uncharacterised: do they encapsulate toxins or form cellular interactions? Or do they do both? This project will use cryo-EM to determine the currently unknown structure of a bacterial teneurin-like protein, and is best suited to someone with a strong interest in protein structure and function.
Te Waiora (The Life-giving waters). A Kaupapa Māori investigation of the biological and chemical characteristics of groundwater as indicators of mauri
Project code: SCI018
Supervisor:
Prof Cate Macinnis-Ng
Sarah Rewi
Discipline: School of Biological Sciences
Project description
With interests in mātauranga-based science research on the rise, it is important these forms of research are responsive to Māori community needs. Understanding the impact of land-use, particularly agricultural activity, on groundwater resources is of key concern to Māori. This project will involve field-based work and data analysis researching into spatial patterns of groundwater chemical composition and microbial communities. It will examine how scientific indicators can assist mana whenua in their assessment of the state of the water’s mauri. No specific skills are required but it is recommended that the candidate has an interest in the interface between mātauranga and science. It is a requirement that the student has whakapapa Māori.
Structural phylogenetics of the bacterial flagellum: FliI, F1Fo-ATP synthetase, and related proteins
Project code: SCI019
Supervisor:
Nicholas J. Matzke
Caroline Puente-Lelievre
Discipline: School of Biological Sciences
Project description
Our lab has recently invented new methods that add protein structure characters to traditional amino acid sequence datasets. This method should improve accuracy and resolution for inferring very deep phylogenetic histories, but more testing is needed. One example system we are working with is the bacterial flagellum, a complex motility system with 25+ interacting proteins that may date back to the Last Common Bacterial Ancestor. One of the most conserved flagellum proteins is the ATPase FliI, so with a structure-informed phylogeny we can readdress classic debates about the relationship between FliI, the homologous F1Fo-synthetase alpha/beta subunits (used to attempt to root the Tree of Life), and related proteins.
The ideal student would have an interest in phylogenetics and/or protein structure (and ideally some coursework, or at least some computer aptitude). Students will learn our structural phylogeny pipeline (using AlphaFold, and Foldseek’s 3Di characters) and associated bioinformatics and phylogenetics skills (e.g. IQtree), and compare results to lab members working on other flagellar proteins.
Structural phylogenetics of the bacterial flagellum: FliF, sporulation proteins, and relatives
Project code: SCI020
Supervisor:
Nicholas J. Matzke
Caroline Puente-Lelievre
Discipline: School of Biological Sciences
Project description
Our lab has recently invented new methods that add protein structure characters to traditional amino acid sequence datasets. This method should improve accuracy and resolution for inferring very deep phylogenetic histories, but more testing is needed. One example system we are working with is the bacterial flagellum, a complex motility system with 25+ interacting proteins that may date back to the Last Common Bacterial Ancestor. The structural base of the flagellum, and perhaps the oldest core of the system, is the MS-ring FliF protein in the inner membrane. It is related in part to the sporulation protein SpoIIIAG, and a structural phylogenetics approach may allow us to better trace early steps in the evolution of the system.
The ideal student would have an interest in phylogenetics and/or protein structure (and ideally some coursework, or at least some computer aptitude). Students will learn our structural phylogeny pipeline (using AlphaFold, and Foldseek’s 3Di characters) and associated bioinformatics and phylogenetics skills (e.g. IQtree), and compare results to lab members working on other flagellar proteins.
How temperature sets the circadian clock on time
Project code: SCI021
Supervisor:
Discipline: School of Biological Sciences
Project description
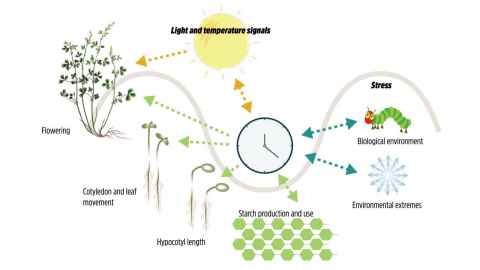
Project: Discover how plants keep time! Like humans and other organisms, plants have an internal clock known as the circadian clock. The clock is set by daily environmental cues, mainly light and temperature cycles happening between day and night. This ensures that key physiological and developmental processes occur at the right time each day, such as plant growth, starch use, responses to stress, and seasonal responses like flowering. While the role of light has been well-studied, little is known about the role of temperature in proper clock functioning. We study the clock in the second most economically important crop group, the legume family, which includes species such as peas, lentils, and lucerne.
The project aims to understand how the circadian clock integrates temperature signals to modulate plant physiology. This knowledge is crucial for breeding climate-resilient legume crops and ensuring food security in the face of climate change.
Skills to be developed: This project will involve the characterization of plants of the model legume species Medicago truncatula in response to temperature fluctuations. It will include genotyping, measuring gene expression, studying plant morphology, and analysing leaf movement rhythmicity as a proxy for circadian clock function.
Ideal candidates (x2): Students with a strong interest in plant molecular biology and plant physiology. BIOSCI 202, 205, 351, and 326 provide a good background.
Figure adapted from Greenham, Kathleen, and C. Robertson McClung. "Integrating circadian dynamics with physiological processes in plants." Nature Reviews Genetics 16.10 (2015): 598-610.
Wounds in microgravity
Project code: SCI022
Supervisor:
Prof Anthony Phillips
Dr Jiwon Hong
Discipline: School of Biological Sciences
Project description
The summer project will provide the motivated candidate with a chance to assist with an existing project developing and testing a device to promote wound healing for use in long distance space flight. The candidate will undertake a mixture of literature searching and summarising as well as assisting with some basic experimental work to evaluate aspects of the technology performance. The project may involve assisting with analysing data or with practical activities like cell culture or imaging as well as other related biological testing activities.
This would best suit a candidate with an interest in space research as well as biology/medicine and/or basic engineering. Some familiarity with Fusion or CAD type software and/or with 3D printing would be a strong advantage but not an absolute prerequisite. The final configuration of activities will be adjusted based on student interests and skills.
Molecular analysis of gene-edited plants for improved flower and fruit development
Project code: SCI023
Supervisor:
Discipline: School of Biological Sciences
Project description
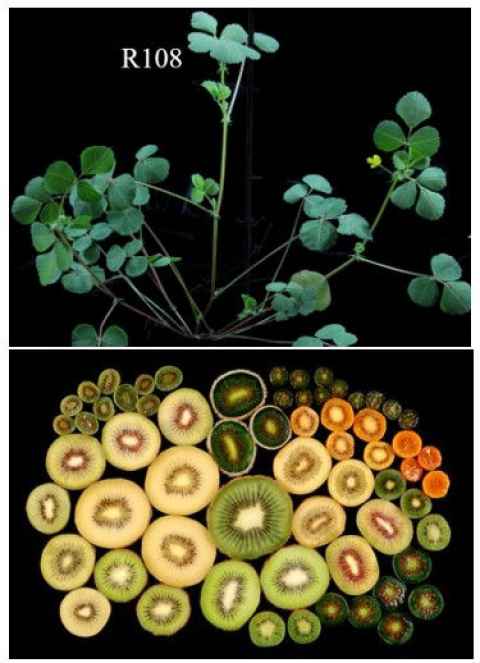
2 projects available. Flowering is an important trait for crops because it affects plant productivity and food security. Legumes are the second most important group of crop plants after the cereals. Plants control flowering differently so each group of plants requires analysis. Thus we study flowering time control in the model legume, Medicago and in kiwifruit (with Plant and Food Research). We are using gene editing to knock out candidate flowering and fruit control genes.
Skill development: You will learn plant molecular biology techniques including how to make gene editing constructs, designing primers, transformation of bacteria and plants and PCR. You will learn how to keep a research lab book, trouble shoot experiments and write up the project report.
Skills required: Skills in molecular biology and genetics with an interest in plant development. BioSci papers such as 202, 351, 355 and 326 provide good background.
Personal attributes: A sense of curiosity. You need to be able to work both as part of a friendly team and independently. To be punctual, work hard and stay focused with strong attention to detail.

The coastal meroplankton of Rangitāhua (the Kermadecs)
Project code: SCI024
Supervisor:
Discipline: School of Biological Sciences
Project description
Plankton samples that have been previously collected at different locations in Rangitāhua will be examined in the laboratory using a microscope, with a focus on quantifying the meroplankton (larval stages of marine invertebrates and fish). Photographic images of rare and abundant meroplankton will be used to contribute to an identification guide for future research at Rangitāhua in a MBIE project (Te Mana o Rangitāhua) with Ngati Kuri and Auckland Museum. An understanding of marine invertebrate diversity would be a distinct advantage (e.g., completion of BIOSCI 208 or similar Invertebrate Diversity course), but on-the-job training will be provided during the project. Ideally suited for a student with interests in biodiversity, identification of small organisms, and in gaining skills in scientific photography and illustration.
Evolving stickiness: Understanding an enzyme-like mechanism that helps to makes bacteria sticky
Project code: SCI025
Supervisor:
Assoc Prof Christopher Squire
Dr. Paul Young
Discipline: School of Biological Sciences
Project description
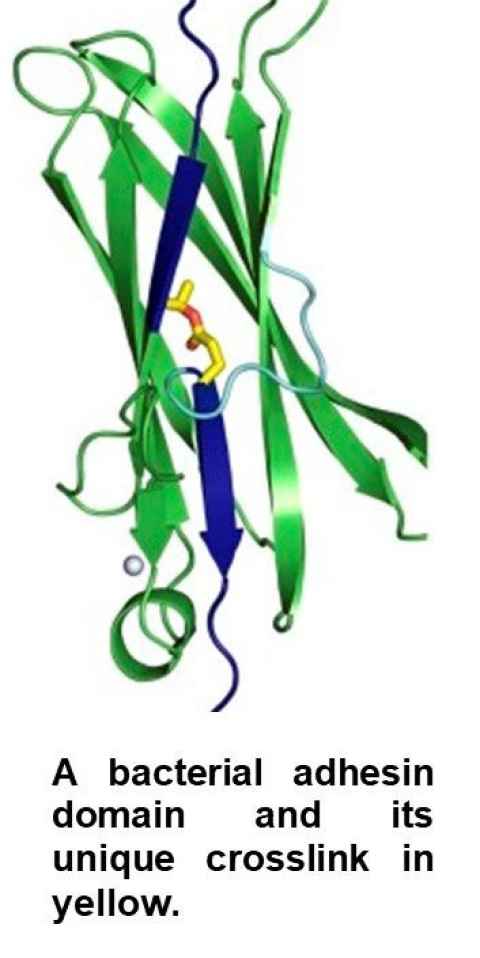
Many Gram-positive bacteria use long narrow adhesin molecules to stick to host cells and to each other in forming biofilms. Adhesins are typically made up of many small protein domains, often containing unique crosslinks to make them ultra-stable (see figure). Some bacteria have evolved a well-known enzymatic mechanism, that of a serine protease, to form the crosslinks – at least that’s what we thought! We have recently discovered some domains that don’t have all the enzymatic machinery (amino acids) we might have expected. You will find out if these proteins still contain crosslinks, and if they do, how you can make the crosslinks form more efficiently by putting the enzymatic machinery back into place – dare we call this a gain-of-function study?
In deconstructing the enzymatic mechanism, we can view these special protein crosslinks through an evolutionary lens. With luck, your summer project results will contribute to a Marsden Fund grant application in 2025 themed on the bacterial evolution of stickiness. Looking to the future, we hope that by understanding exactly how these crosslinks form, we can discover a way to interfere in this process and thus make the lifestyle of the bacterium untenable – it won’t be sticky anymore!
This project will involve cloning, site-directed mutagenesis, protein expression and purification, and characterisation including mass spectrometry fingerprinting. No specific skills are required simply a “can do” attitude and willingness to learn new things.
Targeting protein folding as a novel antibiotic mechanism
Project code: SCI026
Supervisor:
Assoc Prof Christopher Squire
Dr Paul Young
Dr Alan Cameron
Discipline: School of Biological Sciences
Project description
Many Gram-positive bacteria use long narrow adhesin molecules to stick to host cells and to each other in forming biofilms. We want to investigate a potential novel antibiotic mechanism that would interfere with the function of these sticky adhesin molecules.
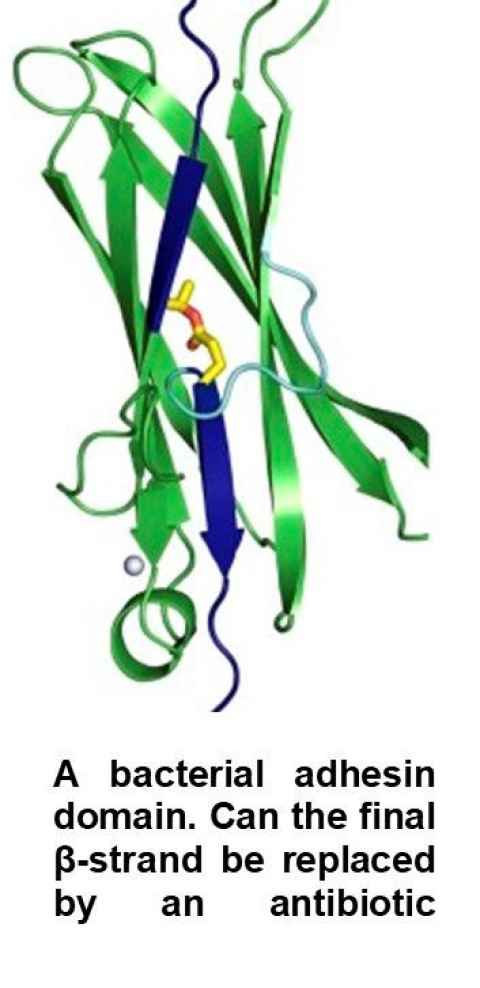
The adhesins are made up of many small protein domains that are secreted from the bacterial cell membrane and then must fold up into their proper 3D shape. In the figure to the right, you can see a blue-coloured peptide – this is the last part of the protein to be completed in folding.
In this project you will learn to propagate Lactococcus lactis, our model bacterium, and then attempt to interfere in its adhesin folding process. As the bacterium secretes its adhesin in your cultures, you will apply a synthetic peptide to try and block the natural folding of the protein. Will this synthetic peptide function as an antibiotic to replace the blue section of the protein (see figure), disrupt proper folding, and thus make the lifestyle of the bacterium untenable?
This project will involve bacterial culture maintenance, biofilm formation and visualisation, and potentially characterisation using fluorescence microscopy, mass spectrometry, and cryoelectron microscopy. You may have the opportunity to produce a synthetic peptide with a fluorescence tag. No specific skills are required simply a “can do” attitude and willingness to learn new things.
Feral cat control: lure effectiveness
Project code: SCI027
Supervisor:
Discipline: School of Biological Sciences
Project description
An effective lure for attracting feral cats to traps and monitoring devices is a major science gap. We will be trialling the effectiveness of novel cat lure compared to conventional lures.
Māori students may also have the opportunity to be involved with a project investigating Māori perspectives on feral cat control (funding-dependent).
Opportunity to work in a larger, supportive team, with researchers from different organisations.

Urban bird distribution and public perceptions
Project code: SCI028
Supervisor:
Discipline: School of Biological Sciences
Project description
This project continues some of our research on urban bird communities and what determines their distribution in cities. The project will include bird counts, some vegetation surveys and some GIS to understand habitat associations and factors that influence bird presence (particularly urban adapters). We also hope to investigate public perceptions of birds, depending on human ethics approval.
Pre-requisites: Must have driver’s licence (preferably full) and access to a car. Must enjoy watching birds! We also hope to survey people about their attitudes to birds, so good communication skills are required.

Exploring Protein–Chemotherapeutic Agent Interactions
Project code: SCI029
Supervisor:
Discipline: School of Biological Sciences
Project description
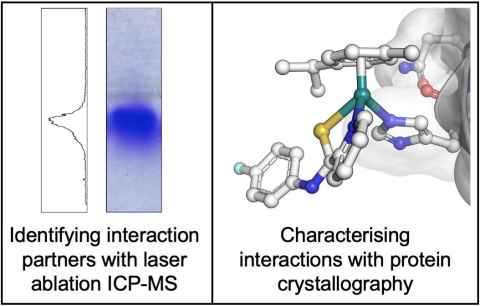
A novel antiproliferative agent plecstatin-1 has been shown to selectively target the protein plectin a known regulator of malignant phenotypes. Plectin is a crucial cytoskeletal component, which acts as a cytolinker. My research project explores the interaction between plecstatin-1 and plectin to understand how selectivity is derived and to understand the role that plectin has within the cytoskeleton. The bioanalytical methodologies surface plasmon resonance and a variety of mass spectrometry techniques (example shown on the left side of the image) will be used to identify the binding domains and regions involved in the plectin–plecstatin-1 interaction. Through protein crystallography (example shown on the right side of the image) and specific binding assays, detailed binding mechanisms and affinities governing the plectin–plecstatin-1 interaction and how this is interrupts the cytoskeleton will be uncovered. The overarching goal is to elucidate how plecstatin-1 interacts with plectin, which in turn interrupts the cytoskeleton and thereby cancer cell behaviour providing crucial insights into the molecular mechanisms driving cancer progression. Ultimately, this deeper understanding holds potential for the development of targeted cancer therapeutics, addressing an unmet need in clinical practice.
Ideal student: No specific skills are required but this project would best suit to someone with an interest in proteins and how they interact with therapeutic agents.
Exploring the functional consequences of different molecular mechanisms in bacterial carbon catabolite repression using mathematical modelling
Project code: SCI030
Supervisor:
Assoc Prof Xue-Xian Zhang
Discipline: School of Biological Sciences
Project description
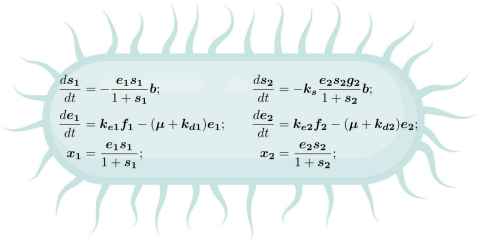
Carbon catabolite repression is a mechanism employed by bacteria to optimize their growth by repressing the expression of genes required for utilising less preferable carbon sources when more preferable ones are available and, thereby, efficiently allocating limited resources for enzyme production. Recent studies have revealed interesting variations in the molecular mechanisms involved in carbon catabolite repression among different bacterial species. For example, carbon catabolite repression in Escherichia coli involves transcriptional regulation, while in Pseudomonas fluorescens, it involves post-transcriptional regulation. This project aims to investigate why such variations exist by identifying the potential factors that have shaped the diversity of molecular mechanisms involved in carbon catabolite repression. The student will develop and analyse mathematical models to compare the dynamics of carbon catabolite repression employing different molecular mechanisms under various environmental conditions. The ideal candidate should have a strong background in molecular biology, a solid understanding of calculus, and proficiency in Python programming. This project offers an opportunity for a highly motivated student to apply their interdisciplinary knowledge and skills in molecular biology, mathematics, and programming to address a fundamental question in microbiology.
Understanding receptor signalling to target migraine
Project code: SCI031
Supervisor:
Discipline: School of Biological Sciences
Project description
Many drugs activate specific G protein-coupled receptors (GPCR) at the cell surface, leading to specific biological outcomes. Several GPCRs represent excellent targets for the treatment of migraine and other neurological disorders. This project will investigate the cellular consequences (intracellular signalling) of activating a receptor, believed to be involved in migraine. Plate-based technologies (eg. Alphascreen, HTRF) will be used to explore signalling in cell culture models. Modern therapeutic concepts, including biased agonism, will be addressed.